MICRONAUT-UR | Bacteriological diagnostics in urinary tract infections
Maximum flexibility and reliability for all requirements of the lab routine combined in a single system

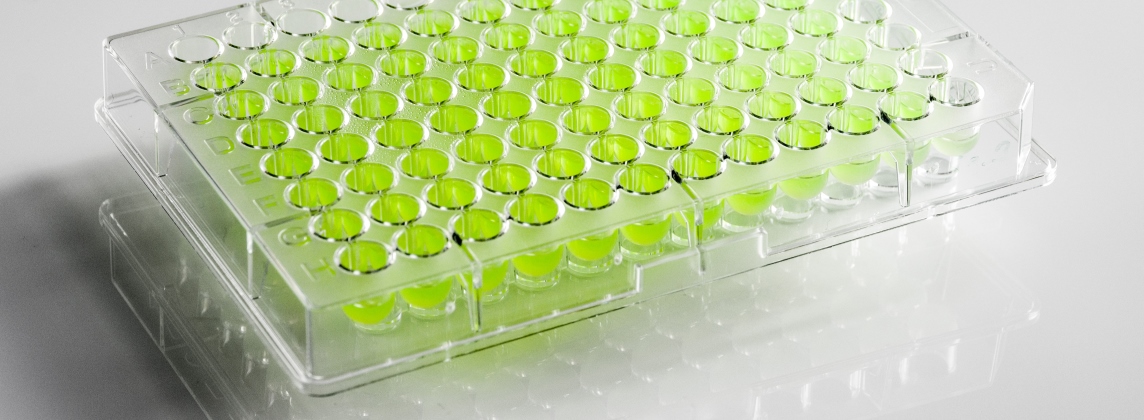
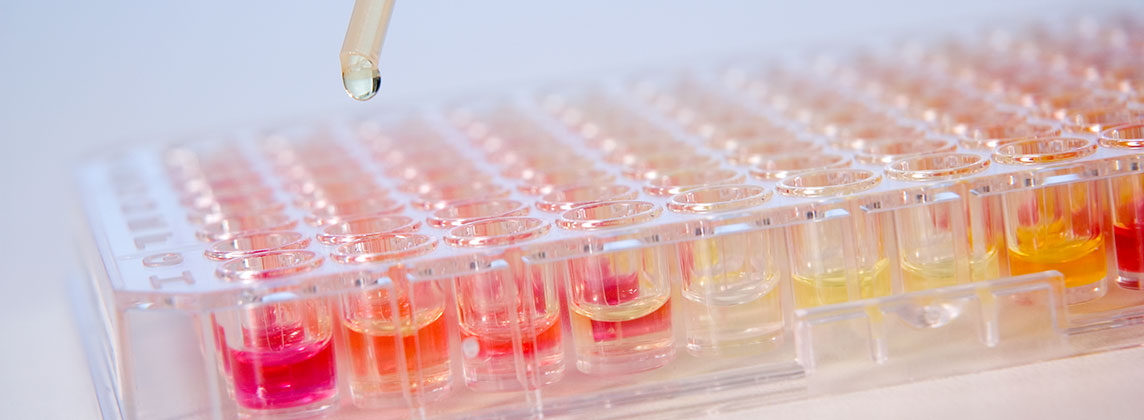
The MICRONAUT-UR system combines biochemical identification (ID) of a total of 54 clinically relevant gram-negative rods and gram-positive cocci with susceptibility testing (AST) of up to 32 UTI-relevant antibiotics using the broth microdilution method in a single working step. Antimicrobial susceptibility testing and interpretation of results are carried out in accordance with the current EUCAST standard.
In addition to AST of antibiotics listed in common UTI guidelines (e.g. ciprofloxacin, levofloxacin, mecillinam, nitrofurantoin, nitroxoline or co-trimoxazole) and susceptibility testing of highly effective reserve antibiotics such as temocillin, vancomycin and meropenem, the MICRONAUT-UR test plate enables the phenotypic detection of resistance mechanisms and phenotypes such as ESBL screening, MRSA detection or the detection of vanA or vanB-mediated glycopeptide resistance in enterococci.
The MICRONAUT-UR system includes a standardized and user-friendly photometer and software-supported evaluation (MICRONAUT software): Plausibility check of AST results by an integrated and up-to-date expert system as well as quality assurance and a statistics module are significantly reducing complexity of result handling and data management using the MICRONAUT software. The possibility of integration into many common lab information systems (LIS) is possible as well.
One isolate can be tested per MICRONAUT-UR test plate - ID and AST combined. Convenient storage at room temperature (15-25°C) and a long shelf life (24 months from date of production) simplify laboratory logistics.
Order directly online
Our sales partner AUROSAN in Essen / Germany is your first contact for these MICRONAUT test products for use in urological diagnostics. You can order the MICRONAUT-UR plates as well as the associated media in their online shop.
The AUROSAN team will also be glad to advise you on whether this MICRONAUT system can be used efficiently both diagnostically and economically. Simply contact them directly >> micronaut@aurosan.de
MICRONAUT ID-Systems
Take here a look at the workflow